10:00 am – 6:50 pm
Tasks
- send info to Stanford Dev Bio.
- finish and submit application to Cornell Physics (due today)
- write to XZ about new experimental design.
- synthesize lots more dsRNA
- start new KD of Ph
- experiment with 2 day KD, 3 day KD, daily serial KD,
- update website
Experiments
- dsRNA production
- ran DCC5 column clean up on PCR products of ph-p and ph-d from last week
- eluted in 20 uL and set up parallel 20 uL T7 reactions with 10 uL product each.
- cell culture
- passaged cells, adherent cells from Schneider’s media culture moved back to SFX — my previous flask of cells in SFX is at too low density.
- cells are ready for more RNAi as soon as I have dsRNA.
- probe production
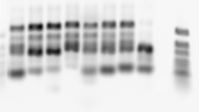
- ran new round of lib4 prep using 1:10 dilution of stock library. Samples C01 to C08 (with reverse primers G01 to G08)
- this amplified sooner (nearing saturation at cycle 24 instead of waiting till after 30).
- still have substantial amounts of large band product in all lanes except well 8. Well 8 was the last one to amplify.
- still see rather high starting values broadly distributed (including 1 unpopulated well). suspect issue with QPCR camera. Should populate a different position on the plate next time.
- also note my old ladder is completely degraded. current ladder on bench degrading a bit.
- embryo labeling
- treated embryo samples 5 min with .1 M HCl (previously skipped in failed labeling last week. BB thinks this step is important for labeling).
- labeled E24 and E20-E24 on embryo sections (previously failed labeling last week).
- incubated 3 min at 90C,
- try staining sectioned embryos with Hb smFISH probes
- using fresh frozen embryo section.
- 1 uL probe in 30 uL hybe dilution buffer, incubating at 37C overnight.
new project planning
- see protected post.
Reading
- Sluis et al 2015 G&D: CRISPR/Cas9 induction of DSBs
- Aymard et al NSMB 2014
- Clynes et al PLoS ONE 2014 (nice immunoFISH)
- Roukos et al, Science 2013
- Iacovoni et al EMBO 2010
