9:30 A – 7:30P
snail imaging
- copy data from confocal (55 GB file). Run O/N at speed 7 using GASP detector
- projecting data from false start. Delete lsm file (mostly blank, and the critical data has been copied into a .mat file). Delete blank images.
- Out of space on local drive D. Moving confocal data 2012 to G: (GRAID) drive.
- # start processing data
Steven lab meeting (see notes)
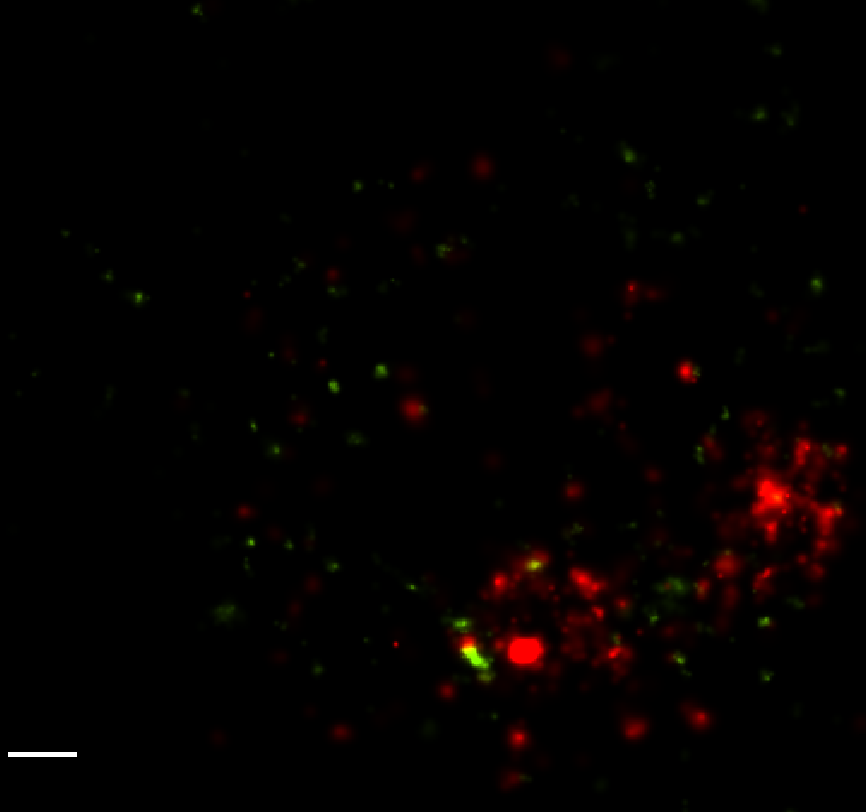
STORM Analysis
- Initial run of dot-fitting complete.
- start writing mini-viewer / interactive viewer for dao-STORM?
Cell culture
- 500 kb probe in situ day 2
- hot wash 15 min, 60C
- appreciable general nuclear background, no detectable specific signal.
- attempting second round of 60C hot wash, see if background cleans up at all.
- 17 min post wash at 60C in SSCT, after brief RT stint in PBT on microscope — background dropped percipitously to nothing. possible weak dots visible in cells.
- Rinse in PBT, block, label nuclear-lamina Cy3B.
Probe making
- Order unlabeled primers for amplification of LC library sequences.
- mixing DNP RNA labeling mix: 250 nmol dUTP, want 3.5 mM solution.
- 250 nmol in 1 mL is 0.250 mM. want 250 nmol in 71.4 uL.
- 7 uL GTP + 7uL CTP + 7 uL ATP + 4.5 uL UTP + 250 nmol DNP-11-UTP, fill to 70 uL.