9:30 A —
Probe making
- pool digests
- aliquot out test DNA: 1 uL digest DNA, 3 uL TBE, 1 uL 6x loading dye + 12x gel green, 5 uL formamide with 5% 5M EDTA
- Pour new 16% urea PAGE gels (2 diagonstic gels and 1 extraction gel). Will need 2 extraction gels if we are going to do all 6 samples. But maybe they won’t all have digested nicely.
- Pre-run gel 1 hr.
- denatured samples at 80C, 5 min. Flush wells. Load Gel. Running 1 hr in waterbath at 52 C.
- column purifying and concentrating DNA samples
- heating fresh elution water to 60C
- CANNOT run PAGE gels with gel-green in loading buffer. DNA doesn’t move, ladder looks all wrong.
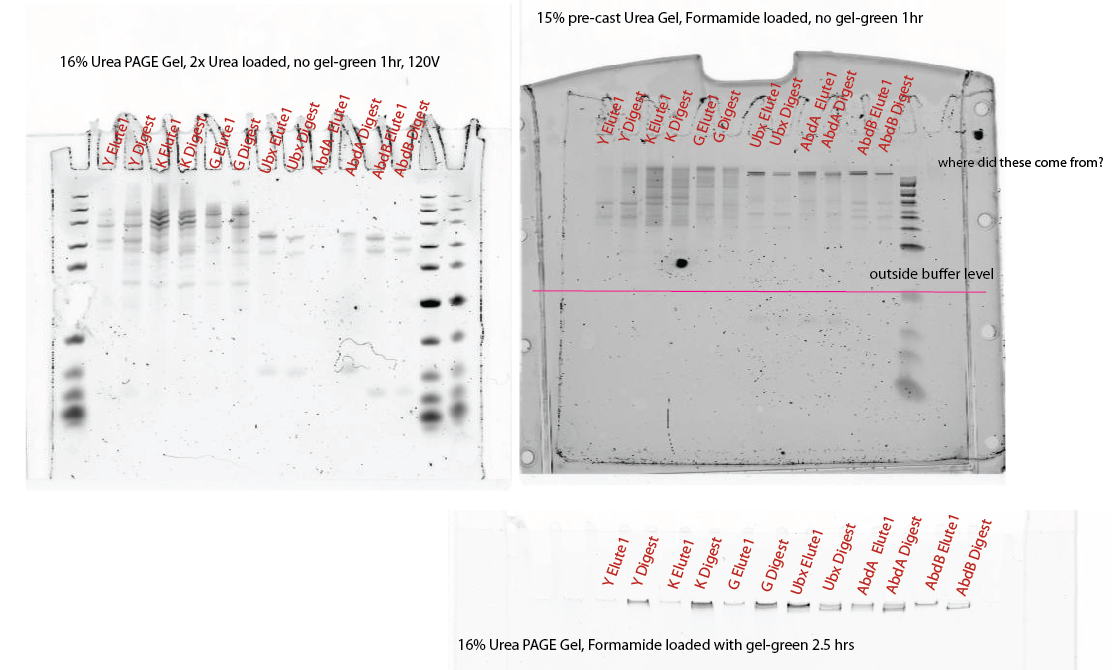
- No change in number of bands before and after nicking.
- If I hadn’t run the pre-digestion DNA, I would say the Ubx, AdbA and AbdB clearly all worked. Previously they gave only 1 band or two bands that were all larger than 100 bp. This is the first time I see a band under 100 bp and the first time I expect a band under 100 bp, but somehow it shows up in the un-nicked elution DNA as well!
- Running another round of 1 mL PCR of Ubx, AdA and AbdB to test on denaturing gel for presence of two bands.
- For Y and G the band at 100 bp in the 16% Urea-loaded gel definetely looks much stronger in the Digest than in the Elute case.
- should probably gel extract these tomorrow anyway.
post-digest elutions
- Y 446. respin 432. 2nd wash eluted with 1st elute: 450
- K 565. respin 580: 2nd wash, elute: 761
- G 336. 378 2nd wash elute2: 380
- AA 107 er. respin 130 er 150 er.
- Ubx 163 er. New column, wash 2, elute with wash1: 380 er.
- AbdB 150 er. respin 150 er. 150
- Y fresh elute 3: 72
- K fresh elute 3: 150
- G fresh elute 3: 90
- Ubx fresh elute 3: 12
- AA new column, fresh elute 3: 30 ugly
- AB new column, fresh elute 3: 12 ugly
STORM
- Brian bringing samples over ~2P.
- running sample 1 O/N. Nuclei clearly visible from 647 channel. Dots also quite bright in conventional image, but not blinking much — not too much switching overall, gave up collection at 40K frames.
STORM analysis
- Looking at en data
- DaoSTORM produces artifacts on dense, non-blinking data — splits into periodic structure.
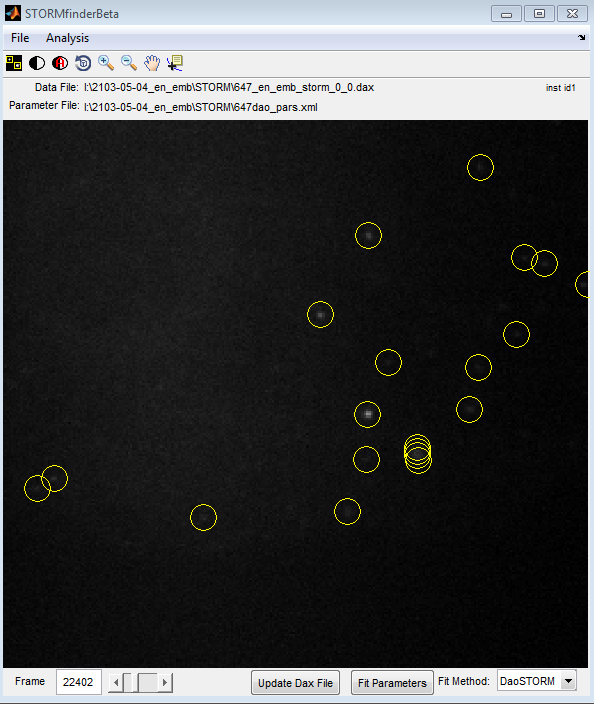
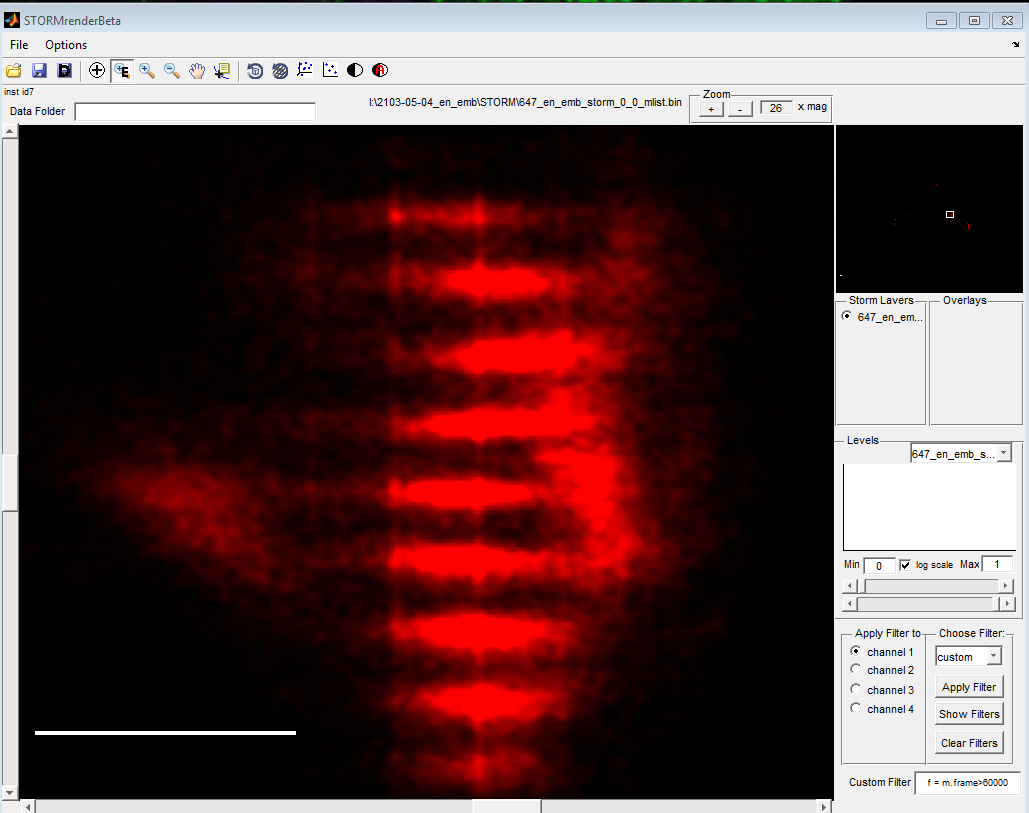
* Raised threshold, test frames look like better fits.
* Re-running on Cajal O/N
Literature
- Reading Mouse hox spatial regulation review (van Steensel 2012)
Banff workshop organizing
- Confirmed participant list so far
- Official BIRS invites sent out.
- 4-7 more open spaces depending on the maybes
- # send out reminder emails
- # send out new invitations
