11:10 A – 7:30 P, 9:30P – 10:30 P
probe making
Labeling amine-tagged probe with cy5 dye.
- labeling reaction for secondary cy5 (see protocol)
- labeled precipitated DNA pellet awaiting HPLC purification (need directions from Hao).
Test large scale reactions with improved nick sites
- take 2 uL in 10 uL TBE for diagnostics
- run 5 uL of this mix on denaturing gel along with aliquot of each sample from before digestion
- ethanol precipitate remaining mix (more glycogen arrived today. Using 3.5 uL glycogen per 200 uL of digestion reaction + 20 uL NaOAc + 700 uL EtOH.
- run RNA probes on gel as well: (~700 and 1800 ng/uL in 100 uL hybe buffer).
Gel = wtf
- sample order: ladder low conc, blue, blue nicked, yellow, yellow nicked, black, black nicked, ubx, ubx nicked, ladder, P1-RNA, P1-rc-RNA, ladder
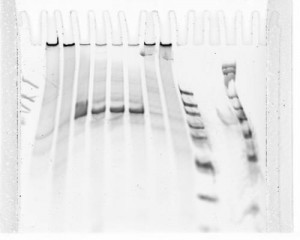
- some samples trapped in wells? why? Both original DNA and nicked DNA?
- Off target bands. What? Did not see at all on small scale. No sign of nicking.
- used ~week old home-cast gel. Try again tomorrow, get the gel to run cleaner. Use percipitated nicked samples.
Hybe tests
- check samples
- 42C 20 min wash in 2X SSCT
- move samples to 4C
- finish checking samples. stains look strong for AATAT-cy5, AATAT-P1 + S1short-cy5 and AATAT-P2 + S2
- en-cy5 seems to have stained as well. Too many dots per cell (~2-6?)
- BX-C not looking too good for staining. This probe had completely evaporated and left a pellet that very seriously did not want to go back into hybe solution despite extensive vortex.
- no primary control has extremely little signal. suggests most of the background is in fact primary mediated?
- heating large hybe oven to 60C for 2x wash in 2x SSCT
Cell labeling, test new probes with 3D FISH protocol
- Split Kc167 cells, passage into new flasks since we are using more.
- Use newest vial of Kc167 cells to coat 6 new coverslips
project 2
- see summary.
STORM
- some how mounted sample C2 upside down — cells destroyed. Imaging session canceled.
