12:20p – 7:00p, 8:40p – 11:30p
Goals
- reread and edit review
- analyze latest round of BXC data
Probe making
- verify direction of MYcroArray BX-C Hox probes: probe region is “+” strand (antisense)
- T7 is on the nick side primer –> RNA will be “-” strand (sense).
- RT will be back to “+” side (antisense)
- Gels (alway ubx, abdA, abdB R to L).
- PAGE gel ladder is 2uL of ULR
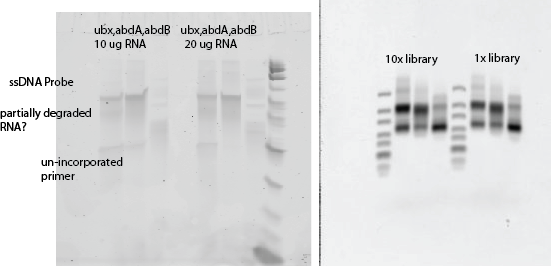
Reverse transcription
- http://tools.invitrogen.com/content/sfs/manuals/superscriptII_pps.pdf
- scaling up to 40 uL reaction.
- in 24 uL, 10 ug of RNA + 300 pmol primer + 2 uL 10mM dNTP
- ~1000 ng/uL -> 1 ug/uL. 10 uL RNA.
- 100 mM primer = 100 pmol/uL = 3 uL primer (use Forward adapter primers). Acutally used 5 for both ubx constructs, see how it compares
- heat to 65C for 5 min. Quench on ice.
- add per reaction (pre-mix this).
- 8uL buffer + 2 uL RNaseOUT/inhibitor + 4 uL .1M DTT
- incubate at 42C 2min
- Add 2 uL (200 units) Superscript II, mix thoroughly by pipetting
- 42C for 50 min+ (ran ~ 2hrs).
- Stop reaction and degrade RNA by alkaline hydrolysis:
* Add 10 uL mix of equal parts 0.5 M EDTA and 1 M NaOH, incubate 70C; 20 minutes
version 2
- try same thing with 20 uL of RNA
- totals (7x): 54 uL bufffer + 14 uL inhibitor + 28 uL DTT. 14uL per reaction
- Running out test gel
- box errors if internal volume not clearly above well (and maybe errors other-times too). Finally got gelbox running.
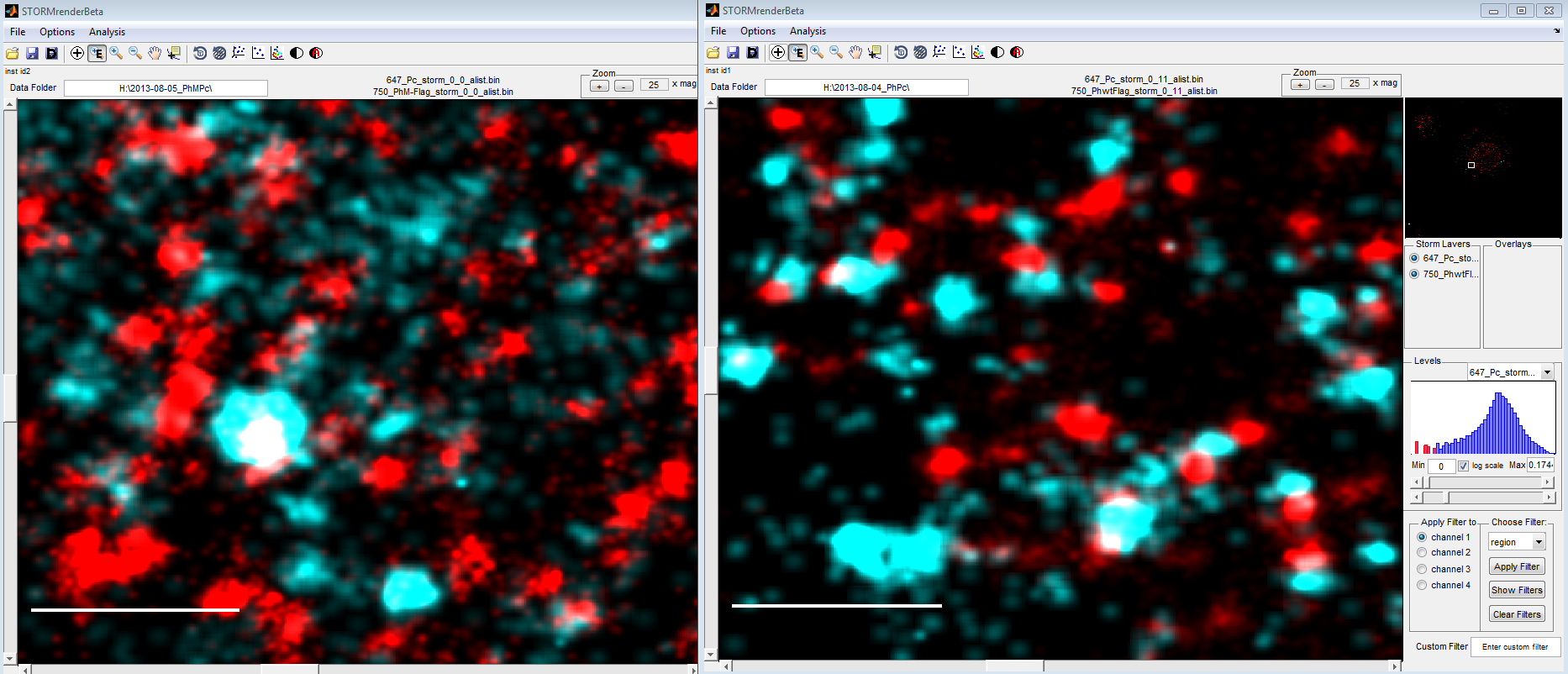
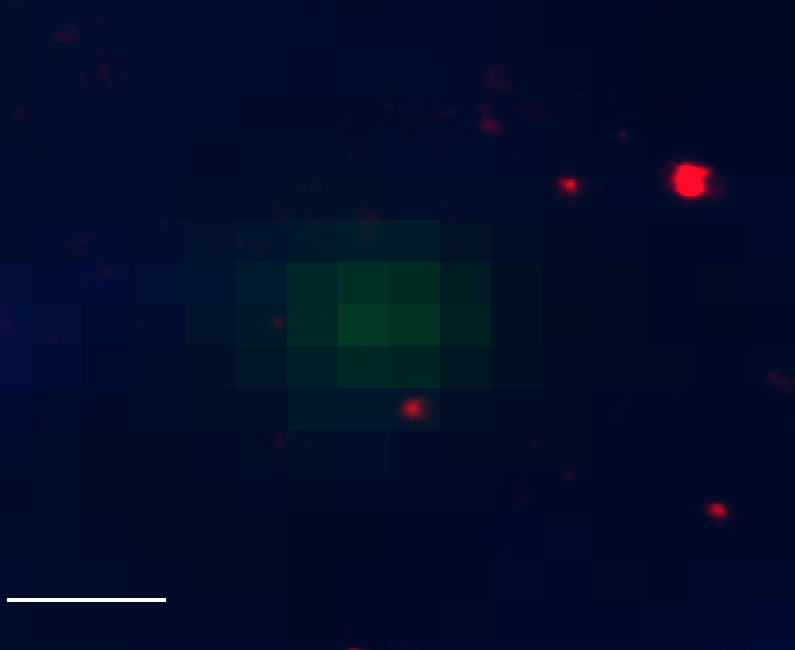
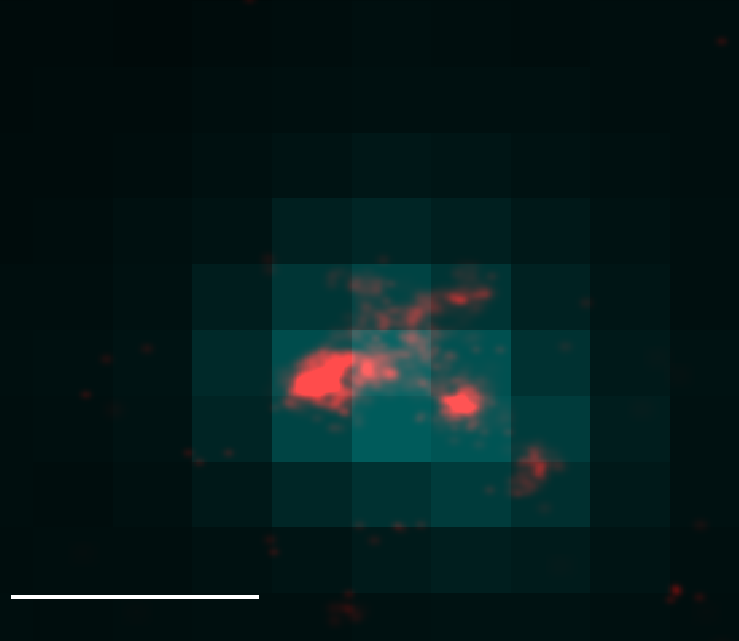
Ph data
antibody labeling
- making A647 and cy7 abcam a-rab and R&D anti-m A647 and cy7
- new aliquiot of cy7 at 2mg/mL — seems less potent than Hao’s aliquots. First attempt with .75 uL of dye gives a bit low labeling.
- attempting again with 2 uL of dye for anti-m (all out of abcam a-rab).
- try labeling same primary in two different secondary colors as a control for co-localization labeling.
Bxc embryo data
- let’s test washout and let’s test yellow locus in embryos.
- we should also start putting beads on the sample again.