9:30a -12:30p
STORM Analysis
- setting up more dot-finding analysis on Cajal
- setting up final splitQVdax for current data.
- Running stv2mat on all Mosaic data not yet processed.
- F10 seems to have issues running a complete round of splitQVdax. Some movies get properly split and some get written out as 0 bytes.
Coding
chromatin simulations for Ph project
- attempting to implement Bond Fluctuation Model based on original published algorithm.
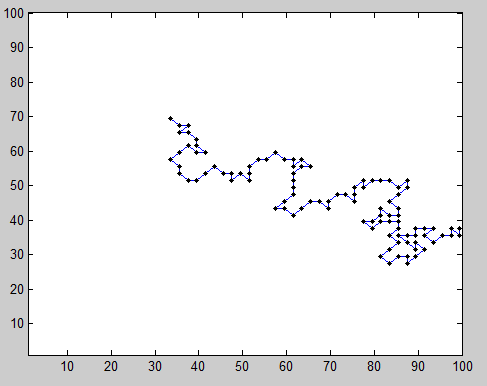
- successfully implemented model in 2D. Should generalize easily to 3D
- code is
BondFlucutationMethod_131014.min\Chromatin\Code\Scripts\ - Adding periodic boundary conditions is difficult because of measuring distances between positions. Might need to switch x,y coordinates to proper spherical coordinates or something.
Next steps
- Attempting to generalize this to Strings and Binders model. (Paper: PNAS)
- SI methods say that all molecules are moved once per MC step. Not clear how this can be implemented, seems a strong departure from the Bond Fluctuation Model.
- Not clear how to add binders and still have polymer be able to move. Maybe bound factors should move as a unit?
- Wrote to senior author for clarification. First author not responding to email.
- Wrote to Ajaz to explain delay.
Working on GUI for dot analysis
- adding flexible user defined parameters
- fixed several dependent functions
- fixed up
startup_demo.m - synched matlab-storm master up with Alistair. Switched back to Alistair branch for
Presentation
- working on slides for Stowers presention.
- Goal1: finish enhancer architecture section.
New hybridizations
- Test F9 sample (from G2, F9, G1 short probes stain). No signal at all. Not good. This worked fine in Bogdan’s set. E2 data may also be background.
- G5-P2-cy3 + S2-cy5 and G4-P1-405 + S1-cy5
- G5-P2-cy3 + S2-cy5 and G4-common-P1 + S-405-dock + T-405
- G5-P1-common + S-405-dock + T-405
