10:00a – 12:50a
Ph Project
- Meeting with Ajaz, discuss paper revisions.
- continue preparing slides for discussion with XZ.
Meeting notes (4p – 7:30p)
To image by Confocal (top priority)
- Endogenous Ph
- Endogenous Pc
- Endogenous Psc (may not work)
- Ph-Flag
- Ph-M-Flag
figure tweaks
- increase contrast in Fig 1 F.
- Thicker scale bars on all panels in Fig 2
- Put more heavily dispersed Ph into Fig 5.
- Fig2 Panel I find example with stronger colocalize
- better Y axis label for histograms.
controls for supplement
- Supplement figure S1.
- No primary controls
- Flag in S2 cells with DAPI contrast.
Fig 5
- change figure back to Ajaz format
- Refer to supplement.
- don’t want to do review experiments to show chromatin template effects
Model for supplement
- Chromatin template important or not?
- Effect of clustering on radius gyration
Expression analysis
- Ajaz will send raw expression files.
- see if we take top 30 Pc target genes, how many increase expression vs. the average gene. Something like this.
Chromatin Project
Sequencing Prep
- Get library from Hao
- check primer concentrations
- 2x Plate PCR of library. D01 – G12
- ran 25 uL scale PCR with T7 and common primers
- pool these, add on T7-NEB adapter in 3 rounds of PCR
Lib2 probes to sequence
- F11-P3 3 replicates of probe synthesis
- F12-P3 2 replicates of probe synthesis
- G09-P1 1 probe sample
- D12-P1 1 probe sample (top left of orange box)
- all diluted 0.5 uL to 200 uL ddH2O
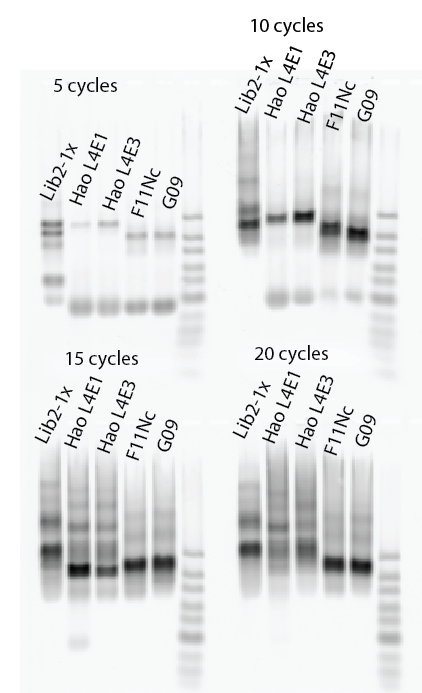
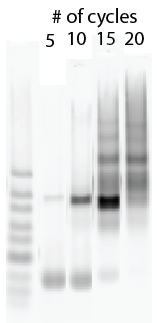
Poor man’s limited cycle PCR:
- Lib2X, Lib4E1, Lib4E3, F11NC, G09
Focus: birth and death of a probe in PCR
LC-PCR
- Lib2-x1 1:32, (2 uL)
- Lib2-x2 1:32,
- Lib2-x1 1:32 v2. (2 uL)
- HaoLib4 E1 (8 uL)
- HaoLib4 E3
- D12-P1 (2 uL)
- G09-P1
- F11-P3 Nc
- Fll-P3 12-18
- F11-P3
- F12 P3 NC
- F12 P3 1580
- Hao L3-uni + L3 i1R
- Hao L3-uni + L3 i2R
- Hao L4 E1F + E1R
- Hao L4 E3F + E3R
Completed limited cycle PCRs and gel extraction and cleanup.