10:00 am – 12:15 am
Chromatin Project
Sequential staining
- snapped coverslip
- tried mounting half on flow-system anyway
- this leaks — not enough pressure seal in the gaskets (especially relative to pump pressure of manual syringe)
- try manual hybes of second broken piece of coverglass
- hybe on glass slide at 47C 30 min
- wash in 50% formamide 2x SSCT at 37C 20 min
- This failed: coverslip upside down 🙁
- try manual hybes with failed stain of the mounted (larger) half of the broken coverglass
- hybe on glass slide at 47C 30 min
- wash in 50% formamide 2x SSCT at 37C 20 min
- Still failed: very weak spots, not all cells have evident spots
- either 30 min is not enough time to hybe or this probe failed (which could explain why the flip failed).
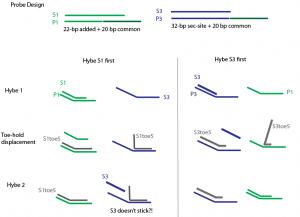
Summary of S1 S3 toehold replacements
New Stains
- Goal: more troubleshooting / testing of sequential staining
- L3E08-p1 + L3E09-p3 + S3-A647
- L3E09-p1 + L3E08-p3
- L3E08-p1 + L3E09-p3
- L3E09-p1 + L3E08-p3
- 4 ul of each probe diluted into 120 uL hybe dilution buffer, split 2 ways (50-60 uL each)
Chromatin Data Analysis
- ChromatinCropper Analysis today
- Finished ChromatinCropper analyzing BXC (last 2 cells)
- ChromatinCropper of F03F04 data — these spots are much smaller than I thought from the quick fit data.
- E06 (repeated for lower background)
- renamed new
BXC_data.matfile asBXC2_data.matand moved to2014-07-09_BlueInternalData - Analyzing G2toG4 data. Looks pretty good. Similar size as ANTC
- launched analysis of E07toE09 data and E07 (rpt 3) data.
- G05 automatic length extraction from name is incorrect — 40 bp, should be 64 !!
- F4, G2, G3, G4 should be double-checked
XZ meeting prep
- Put in draft fig 2.
- bead data for G2toG4 not analyzed, processing now
Simulations
- RC now added openMM to python paths. should try again to see if we can get this to run at all.
- no longer gives error on import simtk.openmm.openmm
- now errors on import dumpOutput
Project 2
- team meeting
- work on code