9:30a – 8:30p
Chromatin labeling
- stains failed again. No stains yet detected with this batch of cells.
- maybe it’s a cell prep issue? This F07 probe worked fine before.
- maybe some reagents went bad in the cell prep? Make new formamide and hybe buffer
Probe Making
qPCR followup
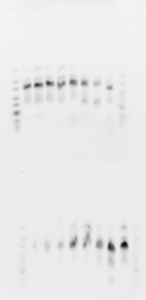
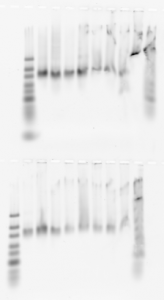
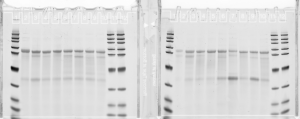
- Run on new 2% gel 2.5 uL of the second elution of DNA from the limited-cycle PCR
- Melted gel. Should remember to always change 100% of the buffer and let’s go back to 10 instead of 11 min.
- lanes: Lib3 E05-E12 (top), F01-F08 (bottom). Also showing yesterdays gel on old gel (not a good week for gels!)
RT reactions
- individual reactions: 10 uL RNA + 4 uL primer + 4 uL RT buffer + 2 uL dNTPs + 1 uL inhibitor + .1 uL
- all prepared with P1 primer
- Master mix: 18*4 uL primer = 72 uL
- 72 uL primer + 72 uL RT buffer + 36 uL dNTPs + 6 uL Maxima enzyme + 18 uL inhibitor. 10 uL each, add to 10 uL RNA.
- out of dNTPs, thawing new dNTPs
- running RT reactions
- using last 2 pre-cast gels. Ordered more gels today.
- also ordered more T7 high yield quick mix kit
- more DCC-5 columns
Troubleshooting Cajal Server 2012 Insight compatability issues
- InsightM gives “Frame out of range!” pop-up error when passed region parameters on Cajal windows 2012.
- This error requires the user to press okay. There-after it seems to analyze just fine. This includes processing the ROI correctly. Batch analysis is suspended until the OK is pressed — very disruptive to automated processing.
- copied files to Monet, run fine using the exact same InsightM version.
- wrote to Bo to comment on bug.
- Meanwhile commented out the ROI parameters on system run in Cajal local.
Cell staining
- passaged cells
- created new 75 cm^2 flask of cells at mid-density for Bogdan.
- plated cells — reprep new cells for hybes. See if a possible cell prep issue is why my hybes suddenly started failing?
- New Hybe Dilution Mix, 10 mL, 1.4X
- 2.8 mL 50% dextran sulfate
- 1.4 mL 2X SSC
- 6.5 mL DI formamide
- 100 uL 10% Tween 20